
═Ō┐Ų╩ųągĄ─▀M▓Įī”╝▒ąį║═┬²ąį╝▓▓ĪĄ─╣▄└ĒŻ¼čėķLē█├³║═▓╗öÓöU┤¾╔·┤µĘČć·Č╝«a╔·┴╦ųž┤¾ė░ĒæĪŻ╚ńłD1╦∙╩ŠŻ¼▀@ą®▀M▓ĮĄ├ęµė┌į\öÓŻ¼│╔Ž±║══Ō┐ŲŲ„ąĄĄ─│ų└m╝╝ąg░lš╣ĪŻ▀@ą®╝╝ągųąŻ¼╔ŅČ╚īW┴Ģī”═ŲäėągŪ░╩ųągęÄäØė╚Ųõųžę¬ĪŻ╩ųągęÄäØųąę¬Ė∙ō■¼FėąĄ─ßt»¤ėøõøüĒėŗäØ╩ųąg│╠ą“Ż¼Č°│╔Ž±ī”ė┌╩ųągĄ─│╔╣”ų┴ĻPųžę¬ĪŻį┌¼FėąĄ─│╔Ž±ĘĮ╩ĮųąŻ¼X╔õŠĆŻ¼CTŻ¼│¼┬Ģ║═MRI╩ŪīŹļHųą│Żė├Ą─ĘĮ╩ĮĪŻ╗∙ė┌ßtīW│╔Ž±Ą─│ŻęÄ╚╬äš░³└©ĮŌŲ╩īWĘųŅÉŻ¼Öz£yŻ¼ĘųĖŅ║═┼õ£╩ĪŻ
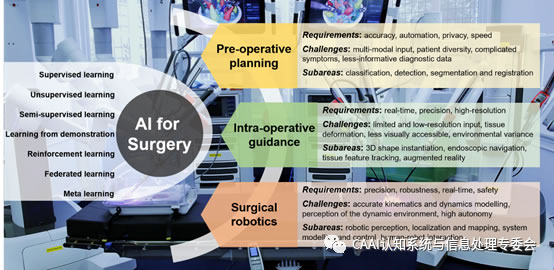
łD1Ż║Ė┼╩÷┴╦┴„ąąĄ─AI╝╝ągŻ¼ęį╝░į┌ągŪ░ęÄäØŻ¼ągųąųĖī¦║══Ō┐Ų╩ųągÖCŲ„╚╦īWųą╩╣ė├Ą─AIĄ─ĻPµIę¬Ū¾Ż¼╠¶æ║═ūėģ^ė“ĪŻ
1ĪóĘųŅÉ
ĘųŅÉ▌ö│÷▌ö╚ļĄ─į\öÓųĄŻ¼įō▌ö╚ļ╩Ūå╬éĆ╗“ę╗ĮMßtīWłDŽ±╗“Ų„╣┘╗“▓Īūā¾włDŽ±ĪŻ│²┴╦é„ĮyĄ─ÖCŲ„īW┴Ģ║═łDŽ±Ęų╬÷╝╝ągŻ¼╗∙ė┌╔ŅČ╚īW┴ĢĄ─ĘĮĘ©š²į┌┼dŲ[1]ĪŻī”ė┌║¾š▀Ż¼ė├ė┌ĘųŅÉĄ─ŠWĮj╝▄śŗė╔ė├ė┌Å─▌ö╚ļīė╠ß╚Īą┼ŽóĄ─ŠĒĘeīė║═ė├ė┌╗žÜwį\öÓųĄĄ─═Ļ╚½▀BĮėīėĮM│╔ĪŻ
└²╚ńŻ¼ėą╚╦╠ß│÷┴╦╩╣ė├GoogleInception║═ResNet╝▄śŗĄ─ĘųŅÉ╣▄Ą└üĒ╝ÜĘųĘ╬░®Ż¼░“ļū░®║═╚ķŽ┘░®Ą─ŅÉą═[2]ĪŻChilamkurthyĄ╚ūC├„╔ŅČ╚īW┴Ģ┐╔ęįūRäe’Bā╚│÷謯¼’B╣Ū╣Ūš█Ż¼ųąŠĆęŲ╬╗║═Ņ^▓┐CTÆ▀├ĶĄ─┘|┴┐ą¦æ¬[3]ĪŻ┼cś╦£╩Ą─┼R┤▓╣żŠ▀ŽÓ▒╚Ż¼┐╔═©▀^裣h╔±ĮøŠWĮjŻ©RNNŻ®īŹĢrŅA£yą─┼K═Ō┐Ų╩ųąg║¾╗╝š▀Ą─╦└═÷┬╩Ż¼─I╦źĮ▀║═ąg║¾│÷č¬[4]ĪŻResNet-50║═Darknet-19ęč▒╗ė├ė┌ī”│¼┬ĢłDŽ±ųąĄ─┴╝ąį╗“É║ąį▓Īūā▀MąąĘųŅÉŻ¼’@╩Š│÷ŽÓ╦ŲĄ─ņ`├¶Č╚║═Ė³GĄ─╠ž«Éąį[5]ĪŻ
2ĪóÖz£y
Öz£y═©│Żęį▀ģĮń┐“╗“Įńś╦Ą─ą╬╩Į╠ß╣®Ėą┼d╚żģ^ė“Ą─┐šķgČ©╬╗Ż¼▓óŪę▀Ć┐╔ęį░³└©łDŽ±╗“ģ^ė“JäeĄ─ĘųŅÉĪŻ═¼śėŻ¼╗∙ė┌╔ŅČ╚īW┴ĢĄ─ĘĮĘ©į┌Öz£yĖ„ĘN«É│Ż╗“ßtīWĀŅørĘĮ├µę▓’@╩Š│÷┴╦ŽŻ═¹ĪŻė├ė┌Öz£yĄ─DCNN═©│Żė╔ė├ė┌╠žš„╠ß╚ĪĄ─ŠĒĘeīė║═ė├ė┌┤_Č©▀ģĮń┐“ī┘ąįĄ─╗žÜwīėĮM│╔ĪŻ
×ķ┴╦Å─4Dš²ļŖūė░l╔õöÓīėÆ▀├ĶŻ©PETŻ®łDŽ±ųąÖz£yŪ░┴ąŽ┘░®Ż¼ī”╔ŅČ╚Čč»BĄ─ŠĒĘeūįäėŠÄ┤aŲ„▀Mąą┴╦ė¢ŠÜŻ¼ęį╠ß╚ĪĮyėŗ║═äė┴”īW╔·╬’īW╠žš„[6]ĪŻī”ė┌Ę╬ĮY╣ØĄ─Öz£yŻ¼╠ß│÷┴╦Š▀ėąą²▐DĘŁūgĮMŠĒĘeŻ©3D G-CNNŻ®Ą─3D CNNŻ¼Š▀ėą┴╝║├Ą─£╩┤_ąįŻ¼ņ`├¶Č╚║═╩šö┐╦┘Č╚[7]ĪŻī”ė┌╚ķŽ┘▓ĪūāĄ─Öz£yŻ¼╗∙ė┌╔ŅČ╚QŠWĮjöUš╣Ą─╔ŅČ╚ÅŖ╗»īW┴ĢŻ©DRLŻ®ė├ė┌Å─äėæBī”▒╚į÷ÅŖMRIųąīW┴Ģ╦č╦„▓▀┬į[8]ĪŻ×ķ┴╦Å─CTÆ▀├ĶųąÖz£y│÷╝▒ąį’Bā╚│÷č¬▓óĖ─╔ŲŠWĮjĄ─┐╔ĮŌßīąįŻ¼LeeĄ╚╚╦[9]╩╣ė├ūóęŌ┴”łD║═Ą³┤·▀^│╠üĒ─ŻĘ┬Ę┼╔õ┐Ųßt╔·Ą─╣żū„┴„│╠ĪŻ
3ĪóĘųĖŅ
ĘųĖŅ┐╔▒╗ęĢ×ķŽ±╦žJ╗“¾w╦žJłDŽ±ĘųŅÉå¢Ņ}ĪŻė╔ė┌įńŲ┌ū„ŲĘųąėŗ╦Ń┘Yį┤Ą─Ž▐ųŲŻ¼├┐éĆłDŽ±╗“ŠĒĘeČ╝▒╗äØĘų×ķąĪ┤░┐┌Ż¼▓óŪęė¢ŠÜ┴╦CNNüĒŅA£y┤░┐┌ųąą─╬╗ų├Ą──┐ś╦ś╦║×ĪŻ═©▀^į┌├▄╝»▓╔śėĄ─łDŽ±┤░┐┌╔Ž▀\ąąCNNĘųŅÉŲ„Ż¼┐╔ęįīŹ¼FłDŽ±╗“¾w╦žĘųĖŅĪŻ└²╚ńŻ¼Deepmedicī”MRIĄ─ČÓ─Ż╩Į─X─[┴÷ĘųĖŅ’@╩Š│÷┴╝║├Ą─ąį─▄[10]ĪŻĄ½╩ŪŻ¼╗∙ė┌╗¼äė┤░┐┌Ą─ĘĮĘ©ą¦┬╩Ą═Ž┬Ż¼ę“×ķį┌įSČÓ┤░┐┌ųž»BĄ─ģ^ė“ųąĢ■ųžÅ═ėŗ╦ŃŠWĮj╣”─▄ĪŻė╔ė┌▀@éĆįŁę“Ż¼╗∙ė┌╗¼äė┤░┐┌Ą─ĘĮĘ©Į³▒╗═Ļ╚½ŠĒĘeŠWĮjŻ©FCNŻ®╚Ī┤·[11]ĪŻĻPµI╦╝Žļ╩Ūė├ŠĒĘeīė║═╔Ž▓╔śėīė╠µōQĘųŅÉŠWĮjųąĄ─╚½▀BĮėīėŻ¼▀@┤¾┤¾╠ßG┴╦ĘųĖŅą¦┬╩ĪŻī”ė┌ßtīWłDŽ±ĘųĖŅŻ¼ųT╚ńU-Net [12][13]ų«ŅÉĄ─ŠÄ┤aŲ„-ĮŌ┤aŲ„ŠWĮjęč’@╩Š│÷┴Ņ╚╦╣─╬ĶĄ─ąį─▄ĪŻŠÄ┤aŲ„Š▀ėąČÓéĆŠĒĘe║═Ž┬▓╔śėīėŻ¼┐╔╠ß╚Ī▓╗═¼▒╚└²Ą─łDŽ±╠žš„ĪŻĮŌ┤aŲ„Š▀ėąŠĒĘe║═╔Ž▓╔śėīėŻ¼┐╔╗ųÅ═╠žš„łDĄ─┐šķgĘų▒µ┬╩Ż¼▓óĮKīŹ¼FŽ±╦ž╗“¾w╦ž├▄╝»ĘųĖŅĪŻį┌[14]ųą┐╔ęįšęĄĮėąĻPė¢ŠÜU-Net▀MąąßtīWłDŽ±ĘųĖŅĄ─▓╗═¼Üwę╗╗»ĘĮĘ©Ą─ŠC╩÷ĪŻ
ī”ė┌ā╚ĖQńRę╚╣▄║═─æĄ└╩ųągųąĄ─ī¦║ĮŻ¼GibsonĄ╚╚╦ [15]╩╣ė├┼“├øĄ─ŠĒĘe║═╚┌║ŽĄ─łDŽ±╠žš„į┌ČÓéĆ│▀Č╚╔ŽĘųĖŅüĒūįCTÆ▀├ĶĄ─Ė╣▓┐Ų„╣┘ĪŻ×ķ┴╦Å─MRI▀Mąą╠ź▒P║═╠źā║┤¾─XĄ─Į╗╗ź╩ĮĘųĖŅŻ¼īóFCN┼cė├æ¶Č©┴xĄ─▀ģĮń┐“║══┐°fĮY║ŽŲüĒŻ¼ŲõųąFCNĄ─║¾ÄūīėĖ∙ō■ė├æ¶▌ö╚ļ▀Mąą┴╦╬óš{[16]ĪŻ╩ųągŲ„ąĄĮńś╦Ą─ĘųĖŅ║═Č©╬╗▒╗Į©─Ż×ķ¤ßłD╗žÜw─Żą═Ż¼▓óŪę╩╣ė├FCNÄū║§īŹĢrĄžĖ·█ÖŲ„ąĄ[17]ĪŻī”ė┌Ę╬ĮY╣ØĘųĖŅŻ¼FengĄ╚═©▀^╩╣ė├║“▀x║Y▀xĘĮĘ©Å─╚§ś╦ėøĄ─Ę╬▓┐CTųąīW┴Ģ▒µäeģ^ė“üĒė¢ŠÜFCNŻ¼ĮŌøQ┴╦ąĶę¬£╩┤_Ą─╩ųäėūóßīĄ─å¢Ņ}[18]ĪŻBaiĄ╚╠ß│÷┴╦ę╗ĘNūį╬ę▒OČĮĄ─īW┴Ģ▓▀┬įŻ¼ęįėąŽ▐Ą─ś╦ėøė¢ŠÜöĄō■üĒ╠ßGU-NetĄ─ą─┼KĘųĖŅŠ½Č╚[19]ĪŻ
4Īó┼õ£╩
┼õ£╩╩Ūā╔éĆßtīWłDŽ±Ż¼¾wĘe╗“─ŻæBų«ķgĄ─┐šķgī”²RŻ¼▀@ī”ė┌ągŪ░║═ągųąęÄäØČ╝╠žäeųžę¬ĪŻé„Įy╦ŃĘ©═©│ŻĄ³┤·Ąžėŗ╦ŃģóöĄ▐DōQŻ¼╝┤ÅŚąįŻ¼┴„¾w╗“BśėŚlŪ·ŠĆ─Żą═Ż¼ęįąĪ╗»ā╔éĆßt»¤▌ö╚ļų«ķgĄ─ĮoČ©Č╚┴┐Ż©╝┤Š∙ĘĮš`▓ŅŻ¼Üwę╗╗»╗źŽÓĻP╗“╗źą┼ŽóŻ®ĪŻĮ³Ż¼╔ŅČ╚╗žÜw─Żą═ęč▒╗ė├üĒ┤·╠µé„ĮyĄ─║─Ģr║═╗∙ė┌ā×╗»Ą─ūóāį╦ŃĘ©ĪŻ
╩Š└²ąįĄ─╗∙ė┌╔ŅČ╚īW┴ĢĄ─┼õ£╩ĘĮĘ©░³└©VoxelMorphŻ¼╦³═©▀^└¹ė├╗∙ė┌CNNĄ─ĮYśŗ║═▌oų·ĘųĖŅüĒīó▌ö╚ļłDŽ±ī”ė│╔õĄĮūāą╬ł÷Ż¼Å─Č°┤¾╗»ś╦£╩łDŽ±Ųź┼õ─┐ś╦║»öĄ[20]ĪŻ╠ß│÷┴╦ę╗éĆė├ė┌3DßtīWłDŽ±┼õ£╩Ą─Č╦ĄĮČ╦╔ŅČ╚īW┴Ģ┐“╝▄Ż¼įō┐“╝▄░³└©╚²éĆļAČ╬Ż║Ę┬╔õūāōQŅA£yŻ¼äė┴┐ėŗ╦Ń║═ĘŪģóöĄ╝Ü╗»Ż¼ęįĮY║ŽĘ┬╔õ┼õ£╩║═╩Ė┴┐äė┴┐ģóöĄ╗»Ą─╣╠Č©╦┘Č╚ł÷[21]ĪŻ╠ß│÷┴╦ę╗ĘNė├ė┌ČÓ─Ż╩ĮłDŽ±┼õ£╩Ą─╚§▒OČĮ┐“╝▄Ż¼įō┐“╝▄ī”Š▀ėą▌^GJäeī”æ¬ĻPŽĄĄ─łDŽ±Ż©╝┤ĮŌŲ╩ś╦║ׯ®▀Mąąė¢ŠÜŻ¼Č°▓╗╩Ūė├ė┌ŅA£y╬╗ęŲł÷Ą─¾w╦žJäe▐DōQ[22]ĪŻ├┐éƱRĀ¢┐ŲĘ“øQ▓▀▀^│╠Č╝ė╔Įø▀^öUÅłĄ─FCNė¢ŠÜĄ─┤·└Ē╔╠▀MąąŻ¼ęį╩╣3D¾wĘe┼c2D X╔õŠĆłDŽ±ī”²R[23]ĪŻRegNet╩Ū═©▀^┐╝æ]ČÓ│▀Č╚▒│Š░Č°╠ß│÷Ą─Ż¼▓óį┌╚╦╣ż╔·│╔Ą─╬╗ęŲ╩Ė┴┐ł÷Ż©DVFŻ®╔Ž▀Mąą┴╦┼Óė¢Ż¼ęįīŹ¼FĘŪäéąį┼õ£╩[24]ĪŻ3DłDŽ±┼õ£╩ę▓┐╔ęį╣½╩Į╗»×ķ▓▀┬įīW┴Ģ▀^│╠Ż¼Ųõųąīó3DįŁ╩╝łDŽ±ū„×ķ▌ö╚ļŻ¼īóŽ┬ę╗éĆ╝čäėū„Ż©╝┤Ž“╔Ž╗“Ž“Ž┬Ż®ū„×ķ▌ö│÷Ż¼▓óīóCNNū„×ķ┤·└Ē[25]ĪŻ
ģó┐╝╬─½IŻ║
[1] G. Litjens, T. Kooi, B. E.Bejnordi, A. A. A. Setio, F. Ciompi, M. Ghafoorian, J. A. Van Der Laak, B. VanGinneken, and C. I. SaĪõnchez, Ī░A survey on deep learning in medical image analysis,Ī▒ Medical Image Analysis, vol. 42, pp. 60©C88, 2017.
[2] P. Khosravi, E. Kazemi, M.Imielinski, O. Elemento, and I. Hajirasouliha, Ī░Deep convolutional neural networks enable discrimination of heterogeneous digital pathology images,Ī▒ EBioMedicine, vol. 27, pp. 317©C328, 2018.
[3] S. Chilamkurthy, R. Ghosh, S.Tanamala, M. Biviji, N. G. Campeau, V. K. Venugopal, V. Mahajan, P. Rao, and P.Warier, Ī░Deep learning algorithms for detection of critical findings in head CT scans: a retrospective study,Ī▒ The Lancet, vol. 392, no. 10162, pp. 2388©C2396,2018.
[4] A. Meyer, D. Zverinski, B.Pfahringer, J. Kempfert, T. Kuehne, S. H. SuĪ¦ndermann, C. Stamm, T. Hofmann, V.Falk, and C. Eickhoff, Ī░Machine learning for real-time prediction of complications in critical care: a retrospective study,Ī▒ The Lancet RespiratoryMedicine, vol. 6, no. 12, pp. 905©C914, 2018.
[5] X. Li, S. Zhang, Q. Zhang, X.Wei, Y. Pan, J. Zhao, X. Xin, C. Qin, X. Wang, J. Li et al., Ī░Diagnosis of thyroid cancer using deep convolutional neural network models applied to sonographic images: a retrospective, multicohort, diagnostic study,Ī▒ The LancetOncology, vol. 20, no. 2, pp. 193©C201, 2019.
[6] E. Rubinstein, M. Salhov, M.Nidam-Leshem, V. White, S. Golan, J. Baniel, H. Bernstine, D. Groshar, and A.Averbuch, Ī░Unsupervised tumor detection in dynamic PET/CT imaging of the prostate,Ī▒ Medical Image Analysis, vol. 55, pp. 27©C40, 2019.
[7] M. Winkels and T. S. Cohen,Ī░Pulmonary nodule detection in CT scan with equivariant CNNs,Ī▒ Medical image analysis, vol. 55, pp. 15©C26, 2019.
[8] G. Maicas, G. Carneiro, A. P.Bradley, J. C. Nascimento, and I. Reid,Ī░Deep reinforcement learning for active breast lesion detection from DCE-MRI,Ī▒ in Proceedings of International Conference on Medical image computing and Computer-Assisted Intervention (MICCAI). Springer, 2017, pp.665©C673.
[9] H. Lee, S. Yune, M. Mansouri,M. Kim, S. H. Tajmir, C. E. Guerrier, S. A. Ebert, S. R. Pomerantz, J. M.Romero, S. Kamalian et al., Ī░An explainable deep-learning algorithm for the detection of acute intracranial hemorrhage from small datasets,Ī▒ NatureBiomedical Engineering, vol. 3, no. 3, p. 173, 2019.
[10]K. Kamnitsas, C. Ledig, V. F.Newcombe, J. P. Simpson, A. D. Kane, D. K. Menon, D. Rueckert, and B. Glocker, Ī░Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation,Ī▒ Medical image analysis, vol. 36, pp. 61©C78, 2017.
[11]J. Long, E. Shelhamer, and T.Darrell, Ī░Fully convolutional networks for semantic segmentation,Ī▒ in proceedings of the IEEE Conference on Computer Vision and Pattern Recognition,2015, pp. 3431©C3440.
[12]O. Ronneberger, P. Fischer, and T. Brox, Ī░U-Net: Convolutional networks for biomedical image segmentation,Ī▒ in Proceedings of International Conference on Medical Image Computing and computer-Assisted Intervention (MICCAI). Springer, 2015, pp. 234©C241.
[13]O. C¸i¸cek, A. Abdulkadir, S.S. Lienkamp, T. Brox, and O. Ronneberger,Ī¦ Ī░3D U-Net: learning dense volumetric segmentation from sparse annotation,Ī▒ in Proceedings of InternationalConference on Medical Image Computing and Computer-Assisted Intervention(MICCAI). Springer, 2016, pp. 424©C432.
[14]X.-Y. Zhou and G.-Z. Yang,Ī░Normalization in training U-Net for 2D biomedical semantic segmentation,Ī▒ IEEERobotics and Automation Letters, vol. 4, no. 2, pp. 1792©C1799, 2019.
[15]E. Gibson, F. Giganti, Y. Hu,E. Bonmati, S. Bandula, K. Gurusamy, B. Davidson, S. P. Pereira, M. J.Clarkson, and D. C. Barratt, Ī░Automatic multi-organ segmentation on abdominal CT with dense networks,Ī▒ IEEE Transactions on Medical Imaging, vol. 37, no. 8,pp.1822©C1834, 2018.
[16]G. Wang, W. Li, M. A. Zuluaga,R. Pratt, P. A. Patel, M. Aertsen, T. Doel, A. L. David, J. Deprest, S.Ourselin et al., Ī░Interactive medical image segmentation using deep learning with image-specific fine-tuning,Ī▒ IEEE Transactions on Medical Imaging, vol.37, no. 7, pp. 1562©C1573, 2018.
[17]I. Laina, N. Rieke, C.Rupprecht, J. P. VizcaĪõıno, A. Eslami, F. Tombari, and N. Navab, Ī░Concurrentsegmentation and localization for tracking of surgical instruments,Ī▒ in Proceedings of International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI).Springer, 2017, pp. 664©C672.
[18]X. Feng, J. Yang, A. F. Laine,and E. D. Angelini, Ī░Discriminative localization in CNNs for weakly-supervised segmentation of pulmonary nodules,Ī▒ in Proceedings of International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI). Springer, 2017,pp. 568©C576.
[19]W. Bai, C. Chen, G. Tarroni,J. Duan, F. Guitton, S. E. Petersen, Y. Guo, P. M. Matthews, and D. Rueckert,Ī░Self-supervised learning for cardiac MR image segmentation by anatomical position prediction,Ī▒ in International Conference on Medical Image Computing and ComputerAssisted Intervention. Springer, 2019, pp. 541©C549.
[20]G. Balakrishnan, A. Zhao, M.R. Sabuncu, J. Guttag, and A. V. Dalca, Ī░VoxelMorph: a learning framework for deformable medical image registration,Ī▒ IEEE Transactions on Medical Imaging,2019.
[21]Z. Shen, X. Han, Z. Xu, and M.Niethammer, Ī░Networks for joint affine and non-parametric image registration,Ī▒in Proceedings of the IEEE Conference on Computer Vision and pattern recognition (CVPR), 2019, pp. 4224©C4233.
[22]Y. Hu, M. Modat, E. Gibson, W.Li, N. Ghavami, E. Bonmati, G. Wang, S. Bandula, C. M. Moore, M. Emberton etal., Ī░Weaklysupervised convolutional neural networks for multimodal image registration,Ī▒ Medical Image Analysis, vol. 49, pp. 1©C13, 2018.
[23]S. Miao, S. Piat, P. Fischer,A. Tuysuzoglu, P. Mewes, T. Mansi, and R. Liao, Ī░Dilated FCN for multi-agent2D/3D medical image registration,Ī▒ in Proceedings of AAAI Conference on artificial intelligence, 2018.
[24]H. Sokooti, B. de Vos, F.Berendsen, B. P. Lelieveldt, I. IĪ”sgum, and M. Staring, Ī░Nonrigid image registration using multi-scale 3D convolutional neural networks,Ī▒ in Proceedings of International Conference on Medical Image Computing and computer-Assisted Intervention (MICCAI). Springer, 2017, pp. 232©C239.
[25]R. Liao, S. Miao, P. deTournemire, S. Grbic, A. Kamen, T. Mansi, and D. Comaniciu, Ī░An artificial agent for robust image registration,Ī▒ in Proceedings of AAAI Conference on Artificial Intelligence, 2017.
 |
| ╔╠ė├ÖCŲ„╚╦ Disinfection Robot š╣ÅdÖCŲ„╚╦ ųŪ─▄└¼╗°šŠ ▌å╩ĮÖCŲ„╚╦Ąū▒P ėŁ┘eÖCŲ„╚╦ ęŲäėÖCŲ„╚╦Ąū▒P ųvĮŌÖCŲ„╚╦ ūŽ═ŌŠĆŽ¹ČŠÖCŲ„╚╦ ┤¾Ų┴ÖCŲ„╚╦ ņF╗»Ž¹ČŠÖCŲ„╚╦ Ę■äšÖCŲ„╚╦Ąū▒P ųŪ─▄╦═▓═ÖCŲ„╚╦ ņF╗»Ž¹ČŠÖC ÖCŲ„╚╦OEM┤·╣żÅS Ž¹ČŠÖCŲ„╚╦┼┼├¹ ųŪ─▄┼õ╦═ÖCŲ„╚╦ łDĢ°^ÖCŲ„╚╦ ī¦ę²ÖCŲ„╚╦ ęŲäėŽ¹ČŠÖCŲ„╚╦ ī¦į\ÖCŲ„╚╦ ėŁ┘eĮė┤²ÖCŲ„╚╦ Ū░┼_ÖCŲ„╚╦ ī¦ė[ÖCŲ„╚╦ ŠŲĄĻ╦═╬’ÖCŲ„╚╦ įŲ█E┐Ų╝╝ØÖÖCŲ„╚╦ įŲ█EŠŲĄĻÖCŲ„╚╦ ųŪ─▄ī¦į\ÖCŲ„╚╦ |